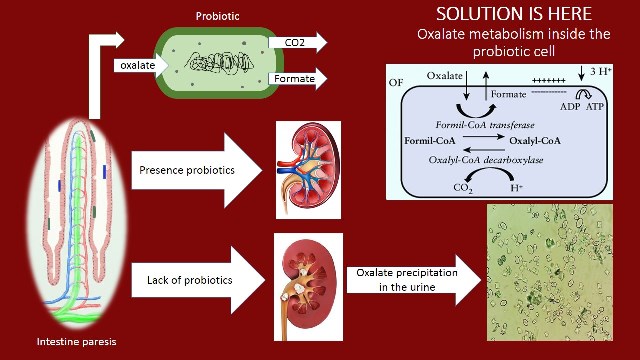
Assessment of the Process Variables for Degradation of Oxalate by Lactobacillus acidophilus ATCC 4356 Using Simulated Rumen Fluid Media and Tea
Applied Food Biotechnology,
Vol. 7 No. 4 (2020),
18 August 2020
,
Page 195-204
https://doi.org/10.22037/afb.v7i4.28600
Abstract
Background and Objective: Lactobacillus acidophilus ATCC 4356 was used for oxalate degradation activity. In the present study, for the first time, the simultaneous influence of process variables on the analysis of high oxalate concentration and its optimization in the simulated intestinal environment was performed. In the end, the optimal results were performed in a tea environment as a common oxalate-containing beverage.
Materials and Methods: After screening the design of ten process variables including pH, glucose, sucrose, inulin, ammonium, sodium oxalate, yeast extract, sodium acetate, inoculum age, and size using Plackett-Burman design, a Box-Behnken method was used with four major variables of pH, glucose, sodium oxalate, and inulin
Results and Conclusion: Results showed that oxalate degradation in simulated rumen fluid was significantly affected by pH and sodium oxalate and glucose concentrations. At optimized conditions, oxalate degradation reached 48.94 ±0.98% of initial concentration. Furthermore, oxalate degradation was investigated in tea (as the most common hot drink in many countries such as Iran) at various times, temperature, and glucose concentration. At optimum condition, oxalate concentration reached 98.86% ±1.05 (from 264 to 24 mg per100 ml).
- Biodegradation
- Lactobacillus acidophilus
- Oxalate
- Plackett-Burman
- Response surface method
How to Cite
References
Morozumi M, Yamaguchi K, Ogawa Y, Kitagawa R. Oxalate metabolism in stone formers. Kidney Dial. 1987; 23: S332-S337.
Miller AW, Choy D, Penniston KL, Lange D. Inhibition of urinary stone disease by a multi-species bacterial network ensures healthy oxalate homeostasis. Kidney Int. 2019; in press.
Karamad D, Khosravi-Darani K, Hosseini H, Tavasoli S, Miller AW. Evaluation of Oxalobacter formigenes DSM 4420 biodegradation activity for high oxalate media content: An in vitro model. Biocatal Agric Biotechnol 2019; 22: 101378.
Abratt VR, Reid SJ. Oxalate degrading bacteria of the human gut as probiotics in the management of kidney stone disease. Adv Appl Microbiol. 2010; 72: 63-87.
Turroni S, Vitali B, Bendazzoli C, Candela M, Gotti R, Federici F. Oxalate consumption by lactobacilli: evaluation of oxalyl‐CoA decarboxylase and formyl‐CoA transferase activity in Lactobacillus acidophilus. Appl Environmen Microbiol. 2007; 76: 5609-5620.
Matthew J, Bull S, Julian P, Marchesi R, Mahenthiralingam, E. The life history of Lactobacillus acidophilus as a probiotic: a tale of revisionary taxonomy misidentification and commercial success. FEMS Microbiol Let. 2013: 349 77-87.
Vilela SF, Barbosa JO, Rossoni RD, Santos JD, Prata MC, Anbinder AL, Jorge AO. Junqueira JC. Lactobacillus acidophilus ATCC 4356 inhibits biofilm formation by C. albicans and attenuates the experimental candidiasis in Galleria mellonella. Virulence. 2015; 6: 29-39.
Kaplan H, Hotkins W. Metabolism of fructooligosaccharides by Lactobacillus paracasei 1195. Appl Environ Microbiol. 2003; 69: 2217-2222.
Ren L, Li G, Jiang D. Characterization of some culture factors affecting oxalate degradation by the mycoparasite Coniothyrium minitans. J Appl Microbiol. 2010; 108: 173–180.
Darilmaz DO, Sonmez S, Beyatli Y. The effects of inulin as a prebiotic supplement and the symbiotic interactions of probiotics to improve oxalate degrading activity. Int J Food Sci Technol. 2018; 54(2): 1-11.
Azcarate-Peril MA, Bruno-Bárcena JM, Hassan HM, Klaenhammer TR. Transcriptional and functional analysis of oxalyl-coenzyme A (CoA) decarboxylase and formyl-CoA transferase genes from Lactobacillus acidophilus. Appl Environ Microbiol. 2006; 72(3): 1891–1899.
Claude P, Champagne H, Gaudreau J, Chartier N. Evaluation of yeast extracts as growth media supplements for lactococci and lactobacilli by using automated spectrophotometry. J Gen Appl Microbiol. 1999; 45(1): 17-21.
Dawson KA, Allison MJ, Hartman PA. Characteristics of anaerobic oxalate-degrading enrichment cultures from the rumen. Appl Environ Microbiol. 1985; 40(4): 840-846.
Campieri C, Campieri M, Bertuzzi V, Swennen E, Matteuzzi D, Stefoni S. Reduction of oxaluria after an oral course of lactic acid bacteria at high concentration. Kidney Int. 2001; 60: 1097-1105.
Murru N, Blaiotta G, Peruzy MF, Santonicola S, Mercogliano R, Aponte M. Screening of oxalate degrading lactic acid bacteria of food origin. Ital J Food Saf. 2017; 6: 6345.
Karamad D, Khosravi-Darani K, Hosseini H, Tavasoli S. Analytical procedures and methods validation for oxalate content estimation. Biointer Res Appl Chem. 2019; 9(5): 4305-4310.
Kargar M, Afkari R, Ghorbani-Dalini S. Oxalate-degrading capacities of gastrointestinal lactic acid bacteria and urinary tract stone formation. Zahedan J Res Med Sci. 2013; 15(10): 54-58.
Tsai JY, Huang JK, Wu TT, Lee YH. Comparison of oxalate content in foods and beverages in Taiwan. JTUA. 2005; 16: 93-99.
Khosravi-Darani K, Zoghi A. Comparison of pretreatment strategies of sugarcane baggase: Experimental design for citric acid production. Biores Technol. 2008; 99: 6986–6993.
Esmaeili S, Khosravi-Darani K, Pourahmad R, Komeili R. An experimental design for production of selenium-enriched yeast. World Appl Sci. 2012; 19:31-37.
Jahadi M, Khosravi-Darani K, Ehsani MR, Mozafari MR, Sabouri AA, Zoghi A. Modeling of proteolysis in Iranian brined cheese using proteinase-loaded nanoliposome. Int J Dairy Technol. 2015; 68: 1-6.
Hadiani MR, Khosravi-Darani K, Rahimifard N, Younesi H. Biosorption of low concentration levels of Lead (II) and Cadmium (II) from aqueous solution by Saccharomyces cerevisiae: Response surface methodology. Biocat Agr Biotechnol. 2018; 15: 25-34.
Zoghi A, Khosravi-Darani K, Sohrabvandi S, Attar H. Patulin removal from synbiotic apple juice using Lactobacillus plantarum ATCC 8014. J Appl Microbiol. 2019; 126(4): 1149-1160.
Asadi SZ, Khosravi-Darani K, Nikoopour H, Bakhoda H. Production of Arachidonic Acid and Eicosapentaenoic Acid by Mortierella alpine CBS 528.72 on Date Waste. Food Technol Biotechnol. 2018; 56(2): 197–207.
Gholami Z, Khosravi-Darani K. An overview of conjugated linoleic acid: microbial production and application. Mini Rev Med Chem. 2014; 14(9): 734-746.
- Abstract Viewed: 705 times
- PDF Downloaded: 592 times